Enzymatic Methyl-seq (EM-seq™)
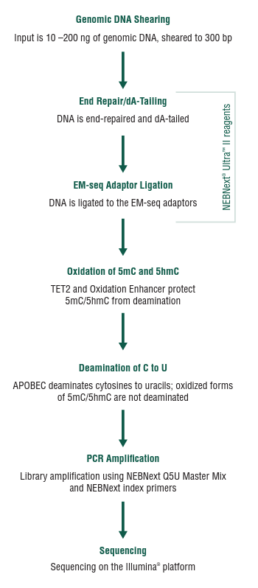
Bisulfite sequencing detects 5-methylcytosine (5mC) and 5-hydroxymethylcytosine (5hmC) at single-base resolution. However, bisulfite treatment damages DNA, which results in fragmentation, DNA loss, and biased sequencing data. To overcome these problems, enzymatic methyl-seq (EM-seq) was developed by NEB, see fig.1 (NEBNext® Enzymatic Methyl-seq Kit, NEB).
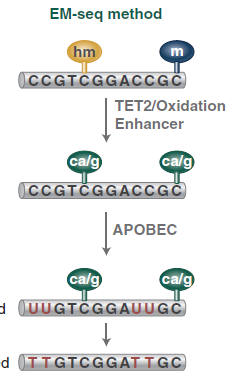
This method detects 5mC and 5hmC using two sets of enzymatic reactions (figure 2).
In the first reaction, TET2 (Ten-eleven-translocation 2 = methylcytosine dioxygenase) and T4-BGT (T4 Phage β-glucosyltransferase) convert 5mC and 5hmC into products that cannot be deaminated by APOBEC3A (Apolipoprotéins B mRNA editing enzyme, catalytic polypeptide-like). In the second reaction, APOBEC3A deaminates unmodified cytosines by converting them to uracils. Therefore, these three enzymes enable the identification of 5mC and 5hmC (Vaisvila et al., 2021).
Platforms to contact for this expertise
Rubriques associées
- Small RNA Sequencing
- Mapping of Transcription Start Sites – TSS
- DNA binding sites map : CUT & RUN
- Chromosome Contact map : 3C, 4C , 5C
- High Chromosome Contact map : HiC-seq
- Mapping of chromatin accessibility sites: DNase seq
- Indirect mapping of chromatin accessibility sites: MNase seq
- Mapping of chromatin accessibility sites: FAIRE seq
- Mapping of chromatin accessibility sites: ATAC seq
- Mapping of RNA-protein interaction sites: CLIP seq
- Mapping of DNA-protein interaction sites: CHIP seq
- Mapping of DNA epigenetic marks: MeDIP
- Mapping of DNA epigenetic marks: Methyl seq
- BiSeq